Note
Click here to download the full example code
1. Quick Start¶
This script produces a default parameter file then uses this to set up a model run. The default parameter file reproduces the constant material properties case in Murphy Quinlan et al., 2021.
import pytesimal.load_plot_save
import pytesimal.quick_workflow
Define a filepath and filename for the parameter file:
folderpath = 'example_default/'
filename = 'example_parameters'
filepath = f'{folderpath}{filename}.txt'
Save a default parameters file to the filepath:
pytesimal.load_plot_save.make_default_param_file(filepath)
You can open this json file in a text editor and change the default values. For this example, we’re just leaving the default values as they are and loading it without editing, and starting a model run:
pytesimal.quick_workflow.workflow(filename, folderpath)
Just wait for a minute or two for your planetesimal to evolve!
# Once 400 millions years has passed and your planetesimal has cooled down,
# we can load the results in to analyse and plot:
filepath = 'example_default/example_parameters_results.npz'
(temperatures,
coretemp,
dT_by_dt,
dT_by_dt_core) = pytesimal.load_plot_save.read_datafile(filepath)
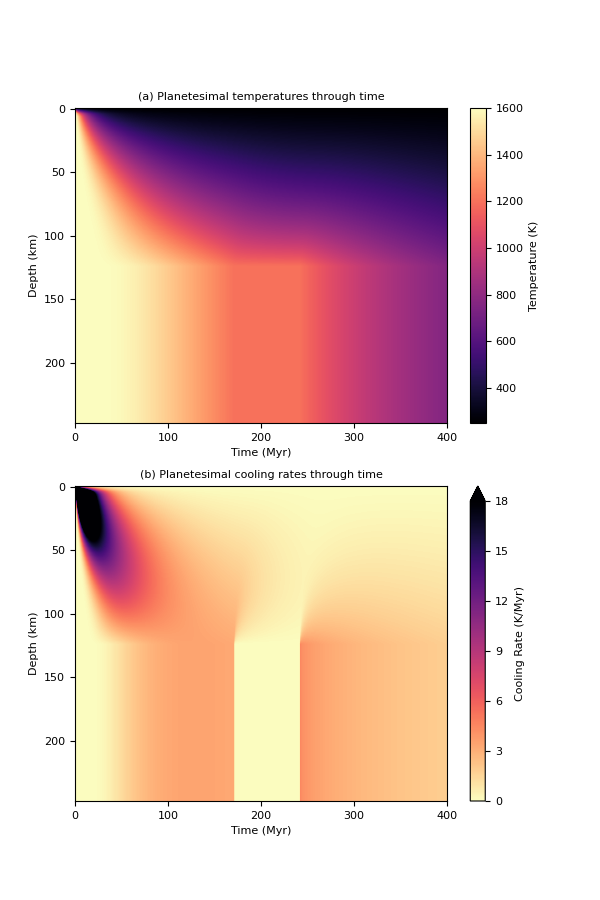
We can visualise the cooling history of the planetesimal:
# Specify a figure width and height
fig_w = 6
fig_h = 9
pytesimal.load_plot_save.two_in_one(fig_w,
fig_h,
temperatures,
coretemp,
dT_by_dt,
dT_by_dt_core)
Total running time of the script: ( 2 minutes 10.762 seconds)